Explore our articles
Using the search function get to know more about us and our products by reading our news, updates and case studies.
Have a support question? Ask it here.
- .com
- 21 CFR Part 11
- 2D-PAGE software
- AI outsourcing
- AI software development
- ATMP
- Automation
- Biopharma
- biotech software development
- biotherapeutics
- Cell and Gene Therapy
- CLIQS
- custom
- custom AI solutions
- Data Integrity
- development
- digital lab transformation
- Digital Transformation
- ELISA automation
- Global Operations
- GMP Compliance
- HCP analysis
- host cell protein
- laboratory AI software
- LC-MS
- life science AI projects
- life science innovation
- life science software
- Life Sciences
- Manufacturing Innovation
- new version
- OEM
- patch notes
- Pharmaceutical Manufacturing
- phoretix 1d
- procurement
- release notes
- reproducibility
- Same Spots
- SAP
- scientist
- software
- Software Solutions
- SpotMap
- SpotMap 2D
- SpotMap ELISA
- SpotMap MS
- supplier
- TotalLab
- TotalLab case studies
- TrendLab
- Vendor Neutral
- Version Control
- Western Blot
- Workflow Automation

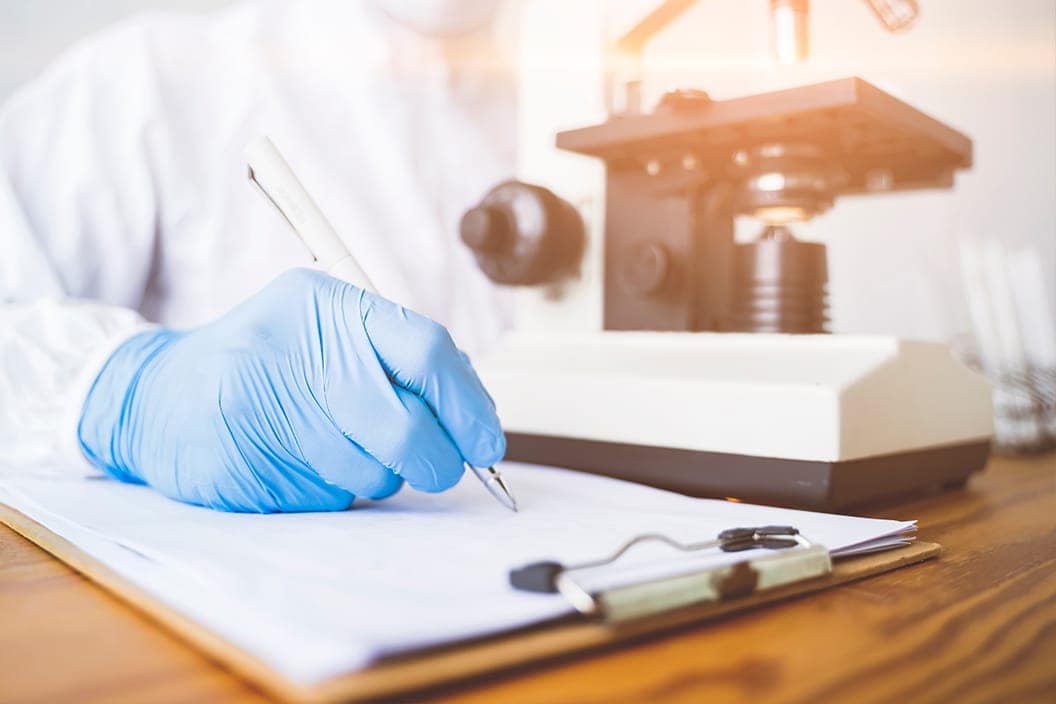
TotalLab’s Dr Steven Dodd Speaks on the Future of Host Cell Protein Analysis in Biotherapeutics
TotalLab automates host cell protein analysis, improving biotherapeutic accuracy, reproducibility, and safety.
Read more >
Unlocking the Power of AI: TotalLab’s Proven AI Software Development for Life Sciences
TotalLab delivers custom AI software for life sciences—explore our case studies and imagine the AI solutions we can create for you.
Read more >
TotalLab Selected to Deliver Custom, Automated ATMP Workflow Solution
TotalLab delivers custom, automated, GMP-compliant ATMP workflow software for global pharma, enabling secure, harmonized analysis.
Read more >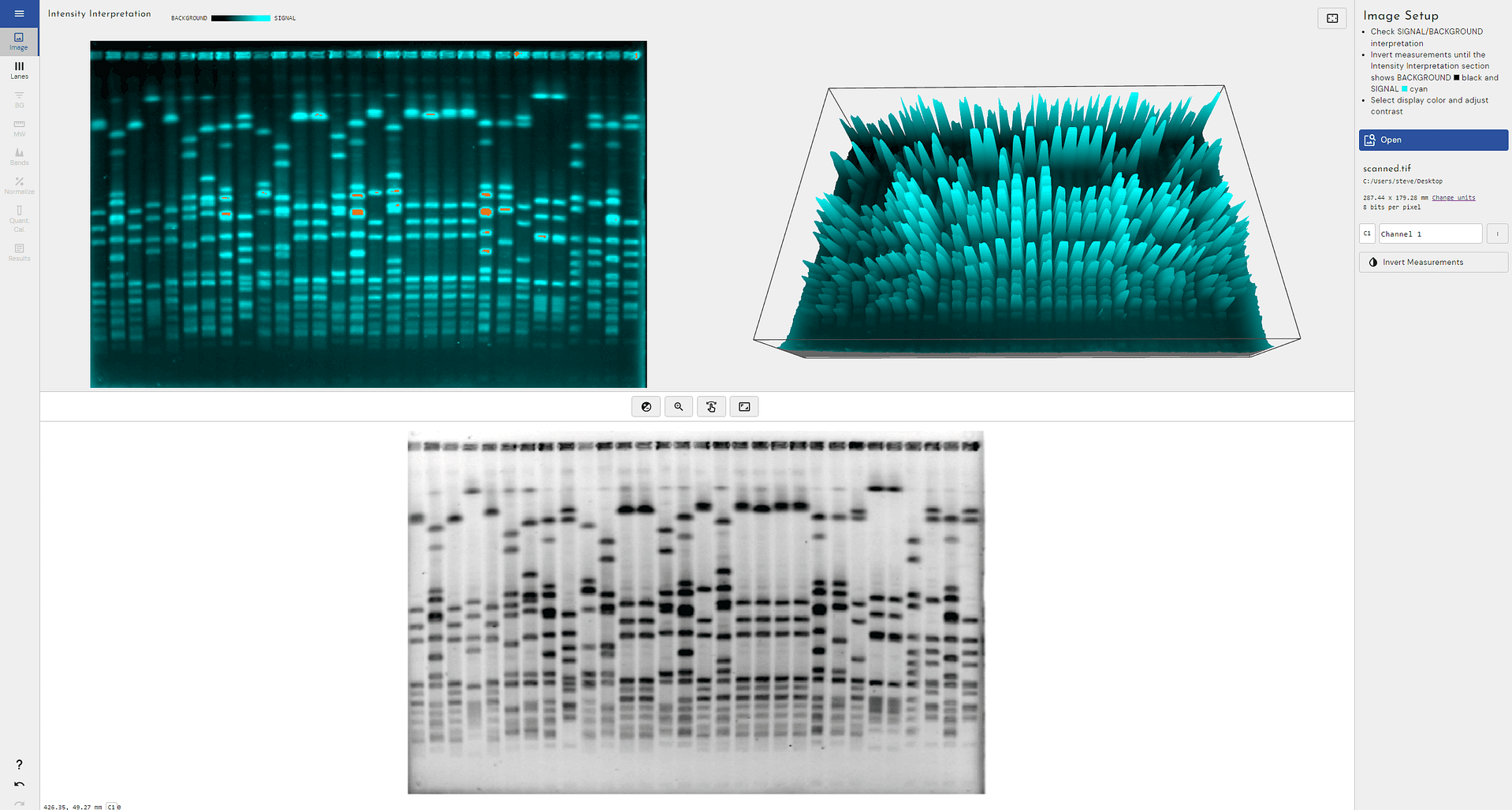
Phoretix 1D 1.3 Has Been Released!
Discover the latest Phoretix 1D software version 1.3—your trusted solution for automated analysis of 1D gels, Western Blots and more
Read more >
What Is Image Analysis Software? Understanding Its Features and Benefits
Read more >

Celebrating 10 Years of TotalLab
In this article we celebrate some of our achievements over the past 10 years and look to the future of TotalLab.
Read more >
Are you using the latest version of SpotMap, our 2D host cell protein image analysis software?
SpotMap users, have you updated to our latest version? If you’re not aware, we’ve recently released version 5.4.578 of SpotMap...
Read more >
Introducing Phoretix 1D
The advanced automation software for fast, reliable, and accurate analysis of 1D gel and blot images.
Read more >
CLIQS – Case Study with the University of Liverpool
Read how we supported a laboratory research team at the University of Liverpool with our 1D gel analysis software.
Read more >
Driving our growth in international markets with two senior appointments
We’re delighted to welcome Dr Steven Dodd as Head of Sales and Business Development and Dave Miller as Head of Technical Services.
Read more >
TotalLab to attend the world’s leading trade fair for lab technology and analysis
We're looking forward to attending Analytica this month in Munich and seeing what innovations are happening in industry...
Read more >
A fresh new look: developing the UI of TotalLab
Read why now was the right time for us to refresh our brand and launch a new look website and what this signals for the future.
Read more >
How to validate the accuracy of your HCP antibody coverage
Discover the importance of host cell protein coverage in biopharmaceutical manufacturing.
Read more >

